Tutorial¶
Here I like to introduce to the usage of the python module pyilt2. If you are just interested in the usage of the command line tool pyilt2report, you can skip this.
the query¶
To start a query on the Ionic Liquids Database you need to use the pyilt2.query() function,
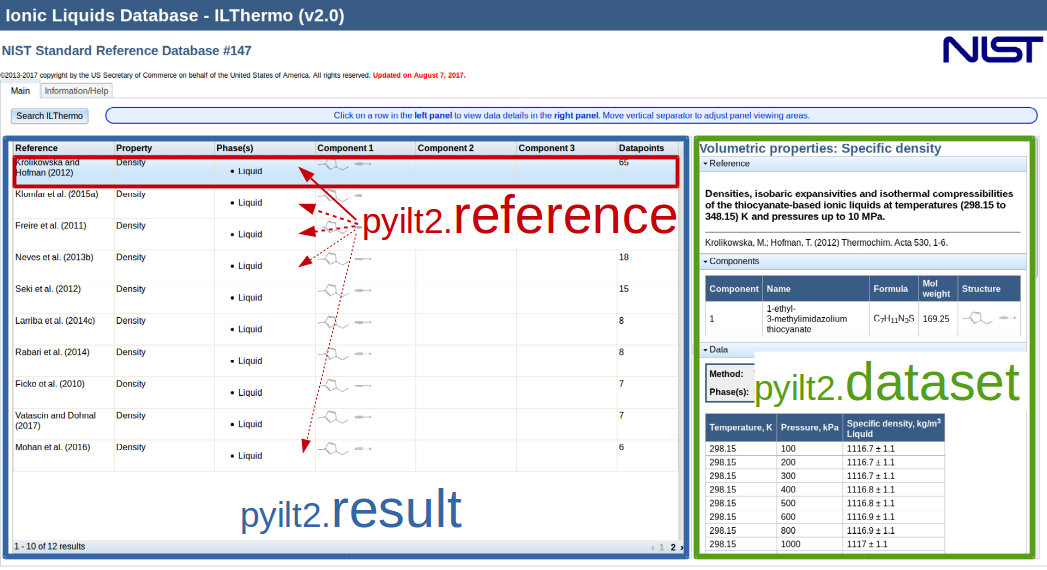
which reproduces the search form (see Fig. 1) from the NIST website.
Each form field is represented by a keyword argument. To specify the physical property you wanna lookup, you have to use the respective abbreviation, as they are listed.
The following code for example will request density data for pure (just one component!) EmimSCN,
and return a pyilt2.result object:
import pyilt2
results = pyilt2.query(comp = "1-ethyl-3-methylimidazolium thiocyanate",
numOfComp = 1,
prop = 'dens')
| keyword | description in search form |
|---|---|
| comp | Chemical formula (case-sensitive), CAS registry number, or name (part or full) |
| numOfComp | Number of mixture components. [Default '0' means any number.] |
| year | Publication year |
| author | Author's last name |
| keywords | Keyword(s) |
| prop | Property [..by abbreviation. Default '' means unspecified.] |
the result & references¶
The pyilt2.result object contanins (like a list) for each hit of the query a pyilt2.reference object.
As list it is iterable, so you can simply iterate over references, like:
>>> # iterate over references
>>> for reference in result:
>>> print (reference)
Krolikowska and Hofman (2012)
Klomfar et al. (2015a)
Freire et al. (2011)
Neves et al. (2013b)
Seki et al. (2012)
Larriba et al. (2014c)
Rabari et al. (2014)
Ficke et al. (2010)
Vatascin and Dohnal (2017)
Mohan et al. (2016)
Klomfar et al. (2015a)
McHale et al. (2008)
As argument in the len() function returns the number of hits,
and of course one can also access the individual references as items:
>>> print ( len(results) )
12
>>> first = results[0]
>>> last = results[-1]
>>> print ('first: {0:s}, {1:s} (data points: {2:d})'.format(first, first.prop, first.np))
first: Krolikowska and Hofman (2012), Density (data points: 65)
>>> print ('last: {0:s}, {1:s} (data points: {2:d})'.format(last, last.prop, last.np))
last: McHale et al. (2008), Density (data points: 1)
The reference object just keeps a few information (see: red box in Fig. 1),
but with the pyilt2.reference.get() method you acquire a pyilt2.dataset object
which finally contains the full data (see: green box in Fig. 1) according to the reference.
>>> dataset = first.get()
>>> print ( type(dataset) )
<class 'pyilt2.dataset'>
the dataset¶
Most important, the property dataset.data contains the data points as
a numpy.ndarray type.
One can access the dimensions of this array by the dataset.shape,
which returns the same as the numpy.shape() function.
>>> print ( type(dataset.data) )
<type 'numpy.ndarray'>
>>> pprint ( dataset.data )
array([[ 2.98150000e+02, 1.00000000e+02, 1.11670000e+03,
1.10000000e+00],
[ 2.98150000e+02, 2.00000000e+02, 1.11670000e+03,
1.10000000e+00],
...
[ 3.38150000e+02, 8.00000000e+03, 1.09590000e+03,
1.10000000e+00],
[ 3.38150000e+02, 1.00000000e+04, 1.09670000e+03,
1.10000000e+00]])
>>> import numpy
>>> print ( numpy.shape(dataset.data) )
(65, 4)
>>> print ( dataset.shape )
(65, 4)
The pyilt2.dataset object owns the following properties...
Note
Sorry.. :( Not finished yet. Please have a look at the API for further details.